Radiation Patterns: Far-Field Body Waves in a Homogeneous Whole Space#
Context: Follow-up lab to the moment tensor module.
Goal: Build intuition for how a moment tensor maps to P and S far-field radiation patterns and how those patterns depend on receiver direction.
Prediction-first rule: For each experiment, write a short prediction before running the cell.
What we model#
Homogeneous, isotropic whole space
Point source at the origin
Far-field only (1/r dependence)
Arbitrary moment tensor \(\mathbf{M}\)
Key references (course): ITS Ch. 9.3 (far-field expressions, nodal structure).
Setup#
We will:
Define a moment tensor \(M_{ij}\) (symmetric 3×3).
Sample takeoff directions \(\gamma\) over the sphere.
Compute:
P radiation amplitude: \(A_P(\gamma) \propto \gamma^T M \gamma\)
S radiation vector: \(\mathbf{A}_S(\gamma) \propto (I - \gamma\gamma^T) M \gamma\)
Visualize:
3D “lobe” plots
Lower-hemisphere equal-area projection
Azimuthal cuts
import numpy as np
import matplotlib.pyplot as plt
# Use consistent figure sizing
plt.rcParams['figure.figsize'] = (7, 6)
plt.rcParams['axes.grid'] = True
Helper functions#
We define:
Spherical grid (theta, phi)
Direction cosines \(\gamma\)
P and S amplitudes for an arbitrary moment tensor
Note: This is geometry-only scaling; we suppress constants like \(1/(4\pi\rho\alpha^3 r)\).
def spherical_grid(n_theta=181, n_phi=361):
theta = np.linspace(0, np.pi, n_theta) # 0..pi
phi = np.linspace(0, 2*np.pi, n_phi) # 0..2pi
TH, PH = np.meshgrid(theta, phi, indexing='ij')
return TH, PH
def gamma_from_theta_phi(theta, phi):
# gamma = [sinθ cosφ, sinθ sinφ, cosθ]
gx = np.sin(theta)*np.cos(phi)
gy = np.sin(theta)*np.sin(phi)
gz = np.cos(theta)
G = np.stack([gx, gy, gz], axis=-1) # (..., 3)
return G
def p_amplitude(M, G):
# A_P = gamma^T M gamma
# einsum: ...i,ij,...j -> ...
return np.einsum('...i,ij,...j->...', G, M, G)
def s_vector(M, G):
# A_S = (I - gg^T) M g
Mg = np.einsum('ij,...j->...i', M, G)
# projection: Mg - g (g·Mg)
gdotMg = np.einsum('...i,...i->...', G, Mg)
return Mg - G * gdotMg[..., None]
def s_amplitude(M, G):
AS = s_vector(M, G)
return np.linalg.norm(AS, axis=-1)
def normalize(M):
# scale moment tensor for display stability
s = np.max(np.abs(M))
return M / s if s != 0 else M
Choose a moment tensor#
Convention (xyz):#
x = East
y = North
z = Up
Start with a simple double couple.
Prediction: Before running, sketch where you expect P nodal planes to be.
# Example 1: a simple DC-like tensor (not tied to strike/dip/rake here)
M = np.array([[0, 1, 0],
[1, 0, 0],
[0, 0, 0]], dtype=float)
M = normalize(M)
M
array([[0., 1., 0.],
[1., 0., 0.],
[0., 0., 0.]])
Compute radiation fields on the sphere#
TH, PH = spherical_grid(n_theta=181, n_phi=361)
G = gamma_from_theta_phi(TH, PH)
AP = p_amplitude(M, G)
AS = s_amplitude(M, G)
AP.min(), AP.max(), AS.min(), AS.max()
(np.float64(-1.0),
np.float64(1.0000000000000002),
np.float64(0.0),
np.float64(1.0))
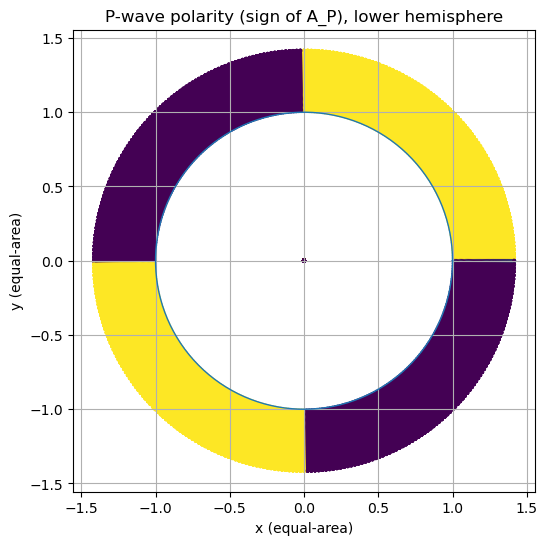
Visualization 1: Lower-hemisphere polarity map (P)#
We plot the sign of \(A_P\) on the lower hemisphere (takeoff directions with \(\gamma_z \le 0\)).
Interpretation prompt:
Identify nodal lines (where \(A_P = 0\)).
How many quadrants do you see?
Which regions are compressional vs. dilatational?
(Recall: in the simplest convention, sign corresponds to first-motion polarity.)
def lower_hemisphere_equal_area(G, A):
# Lambert azimuthal equal-area projection for lower hemisphere
# For lower hemisphere, use points with gz <= 0
gx, gy, gz = G[...,0], G[...,1], G[...,2]
mask = gz <= 0
# colatitude for lower hemisphere: theta in [pi/2, pi]
theta = np.arccos(np.clip(gz, -1, 1))
# radius in equal-area projection: r = sqrt(2) * sin(theta/2)
r = np.sqrt(2) * np.sin(theta/2)
# projected coords
x = r * (gx / np.sqrt(gx**2 + gy**2 + 1e-15))
y = r * (gy / np.sqrt(gx**2 + gy**2 + 1e-15))
return x[mask], y[mask], A[mask]
x, y, a = lower_hemisphere_equal_area(G, AP)
fig, ax = plt.subplots()
sc = ax.scatter(x, y, c=np.sign(a), s=4)
ax.set_aspect('equal', 'box')
ax.set_title("P-wave polarity (sign of A_P), lower hemisphere")
ax.set_xlabel("x (equal-area)")
ax.set_ylabel("y (equal-area)")
# Draw unit circle boundary
t = np.linspace(0, 2*np.pi, 400)
ax.plot(np.cos(t), np.sin(t), linewidth=1)
plt.show()
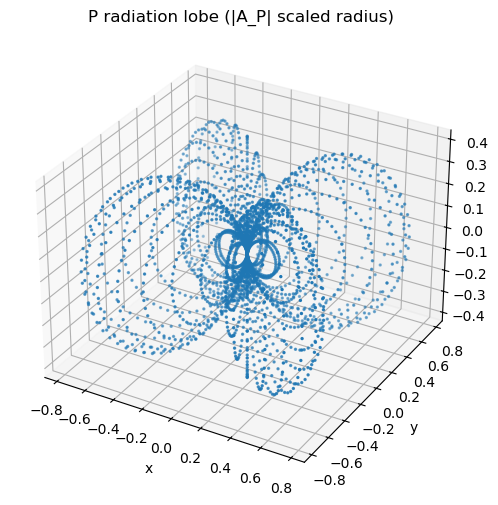
Visualization 2: Radiation lobes (P and S)#
We plot lobe magnitude by scaling the radius with \(|A|\).
Prompt:
P: why are there nodal planes?
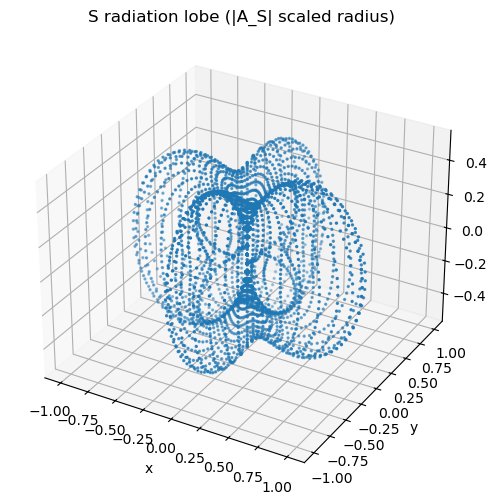
S: why do you expect nodal points (not planes)?
def lobe_xyz(G, A):
# scale direction by |A| (normalized)
An = np.abs(A)
An = An / (An.max() + 1e-15)
X = G[...,0] * An
Y = G[...,1] * An
Z = G[...,2] * An
return X, Y, Z
# Downsample for plotting speed
step_t, step_p = 3, 6
Gd = G[::step_t, ::step_p, :]
APd = AP[::step_t, ::step_p]
ASd = AS[::step_t, ::step_p]
Xp, Yp, Zp = lobe_xyz(Gd, APd)
Xs, Ys, Zs = lobe_xyz(Gd, ASd)
fig = plt.figure(figsize=(7,6))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(Xp.ravel(), Yp.ravel(), Zp.ravel(), s=2)
ax.set_title("P radiation lobe (|A_P| scaled radius)")
ax.set_xlabel("x"); ax.set_ylabel("y"); ax.set_zlabel("z")
plt.show()
fig = plt.figure(figsize=(7,6))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(Xs.ravel(), Ys.ravel(), Zs.ravel(), s=2)
ax.set_title("S radiation lobe (|A_S| scaled radius)")
ax.set_xlabel("x"); ax.set_ylabel("y"); ax.set_zlabel("z")
plt.show()
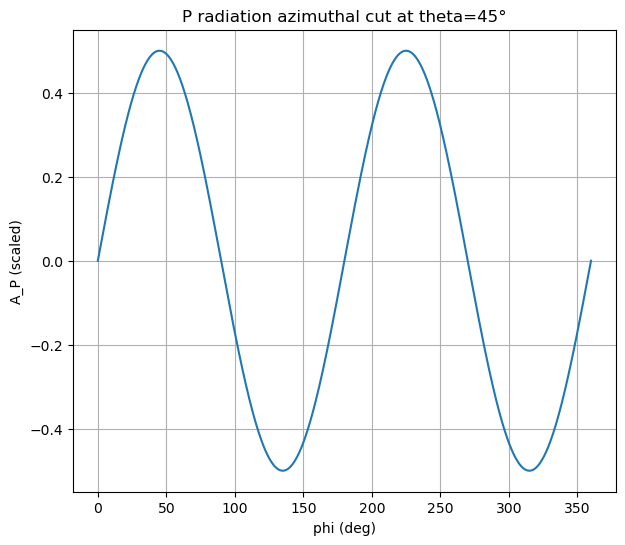
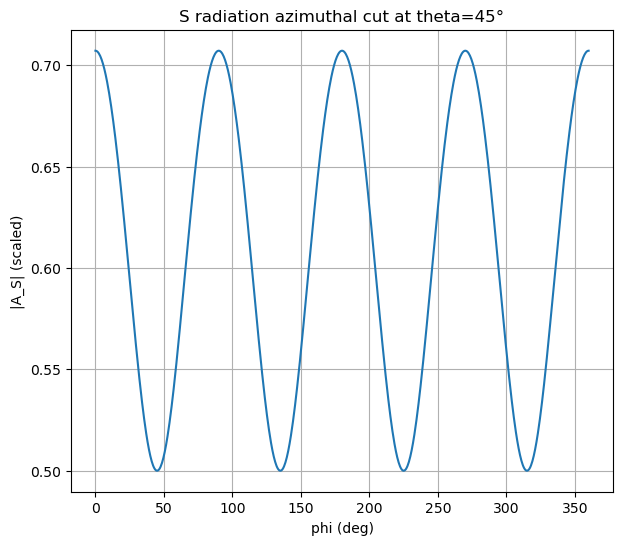
Visualization 3: Azimuthal cut (fixed takeoff angle)#
Fix \(\theta\) and plot \(A_P(\phi)\).
Prediction: What periodicity do you expect for a double-couple-like source?
theta0_deg = 45
theta0 = np.deg2rad(theta0_deg)
phi = np.linspace(0, 2*np.pi, 720)
Gcut = gamma_from_theta_phi(theta0*np.ones_like(phi), phi)
Ap_cut = p_amplitude(M, Gcut)
As_cut = s_amplitude(M, Gcut)
fig, ax = plt.subplots()
ax.plot(np.rad2deg(phi), Ap_cut)
ax.set_title(f"P radiation azimuthal cut at theta={theta0_deg}°")
ax.set_xlabel("phi (deg)")
ax.set_ylabel("A_P (scaled)")
plt.show()
fig, ax = plt.subplots()
ax.plot(np.rad2deg(phi), As_cut)
ax.set_title(f"S radiation azimuthal cut at theta={theta0_deg}°")
ax.set_xlabel("phi (deg)")
ax.set_ylabel("|A_S| (scaled)")
plt.show()
Experiments (Prediction-first)#
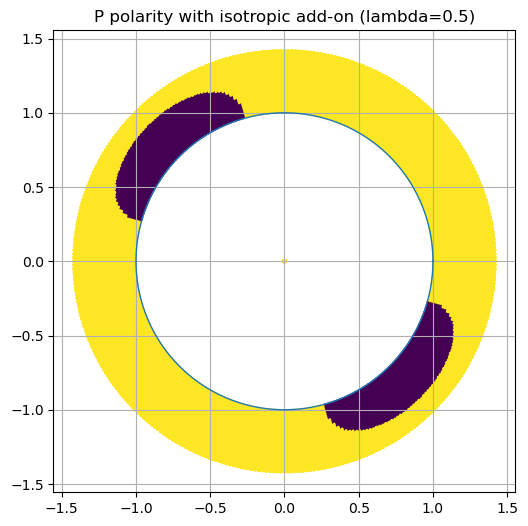
Experiment A — Add isotropic component#
Set \(M \leftarrow M + \lambda I\).
Prediction prompts:
What happens to P nodal planes as \(\lambda\) increases?
Does S change? Why/why not?
lam = 0.5
Mi = normalize(M + lam*np.eye(3))
APi = p_amplitude(Mi, G)
ASi = s_amplitude(Mi, G)
x, y, a = lower_hemisphere_equal_area(G, APi)
fig, ax = plt.subplots()
ax.scatter(x, y, c=np.sign(a), s=4)
ax.set_aspect('equal', 'box')
ax.set_title(f"P polarity with isotropic add-on (lambda={lam})")
t = np.linspace(0, 2*np.pi, 400)
ax.plot(np.cos(t), np.sin(t), linewidth=1)
plt.show()
print("S amplitude range:", ASi.min(), ASi.max())
S amplitude range: 0.0 1.0
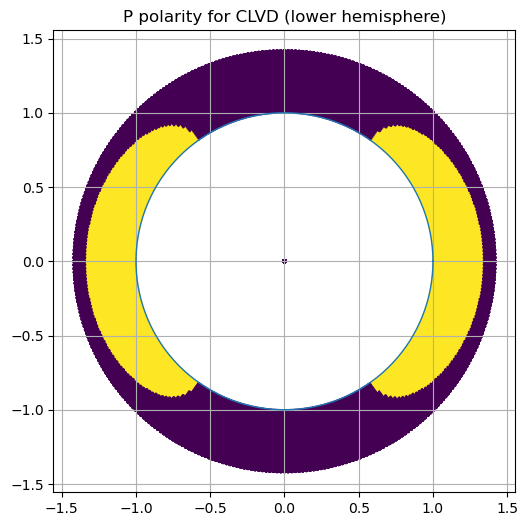
Experiment B — CLVD-like tensor#
Use eigenvalues \([1, -1/2, -1/2]\).
Prediction: How many lobes for P? Where are nodal structures?
Mclvd = np.diag([1.0, -0.5, -0.5])
Mclvd = normalize(Mclvd)
APc = p_amplitude(Mclvd, G)
x, y, a = lower_hemisphere_equal_area(G, APc)
fig, ax = plt.subplots()
ax.scatter(x, y, c=np.sign(a), s=4)
ax.set_aspect('equal', 'box')
ax.set_title("P polarity for CLVD (lower hemisphere)")
t = np.linspace(0, 2*np.pi, 400)
ax.plot(np.cos(t), np.sin(t), linewidth=1)
plt.show()
Experiment C — Random symmetric tensor#
Goal: Practice reading patterns without “knowing the answer.”
Prediction: Before plotting, guess whether the map will resemble DC/CLVD/isotropic.
rng = np.random.default_rng(7)
A = rng.normal(size=(3,3))
Mr = (A + A.T)/2
Mr = normalize(Mr)
APr = p_amplitude(Mr, G)
x, y, a = lower_hemisphere_equal_area(G, APr)
fig, ax = plt.subplots()
ax.scatter(x, y, c=np.sign(a), s=4)
ax.set_aspect('equal', 'box')
ax.set_title("P polarity for random symmetric moment tensor")
t = np.linspace(0, 2*np.pi, 400)
ax.plot(np.cos(t), np.sin(t), linewidth=1)
plt.show()
print("Moment tensor used:", Mr)
Cell In[10], line 17
print("Moment tensor used:
^
SyntaxError: unterminated string literal (detected at line 17)
Lab wrap-up questions#
In your own words: what is \(A_P(\gamma)\) measuring geometrically?
Compare DC vs CLVD: what symmetry difference is most diagnostic?
Why does adding isotropic moment remove nodal planes for P?
For S: why is polarization (direction) unavoidable for interpretation?
Extension (optional):
Implement a simple “first-motion pick” simulator: sample random station directions, label sign(AP), and attempt to recover nodal planes.


